Class 12 : Biology (English) – Lesson 5: Molecular Basis of Inheritance
EXPLANATION & SUMMARY
✨ Introduction
🧬 The molecular basis of inheritance explains how DNA acts as the genetic material, its structure, replication, transcription, translation, regulation of gene expression, and human genome mapping.
🌱 This chapter connects genetics with molecular biology and is key to understanding heredity at the biochemical level.
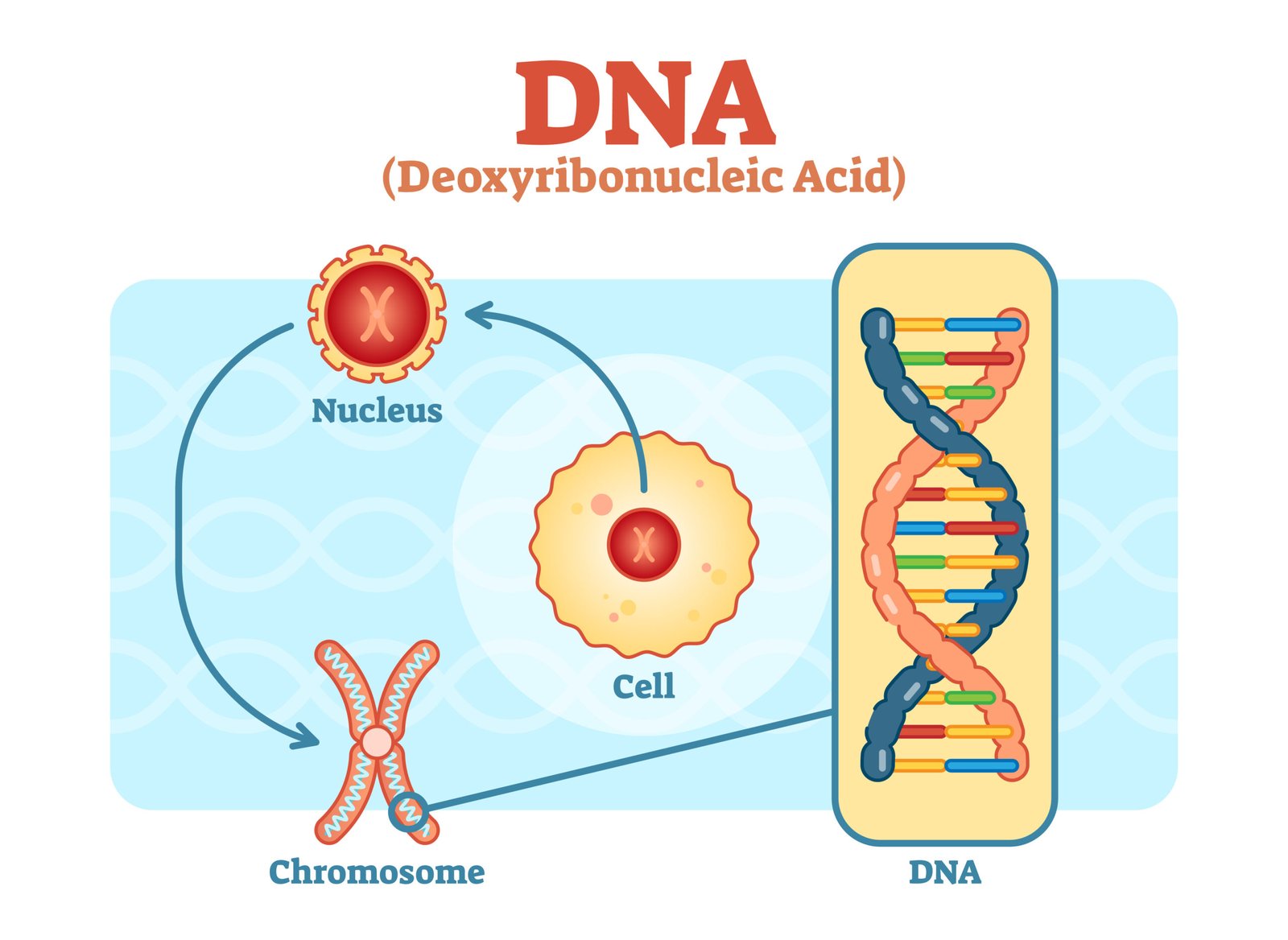
🧬 DNA — The Genetic Material
💡 Griffith’s Experiment (1928): Transformation principle in Streptococcus pneumoniae.
🧪 Avery, MacLeod & McCarty (1944): Proved DNA is the genetic material.
🧪 Hershey–Chase Experiment (1952): Blender experiment with T2 phage confirmed DNA is genetic material.
🧬 Structure of DNA
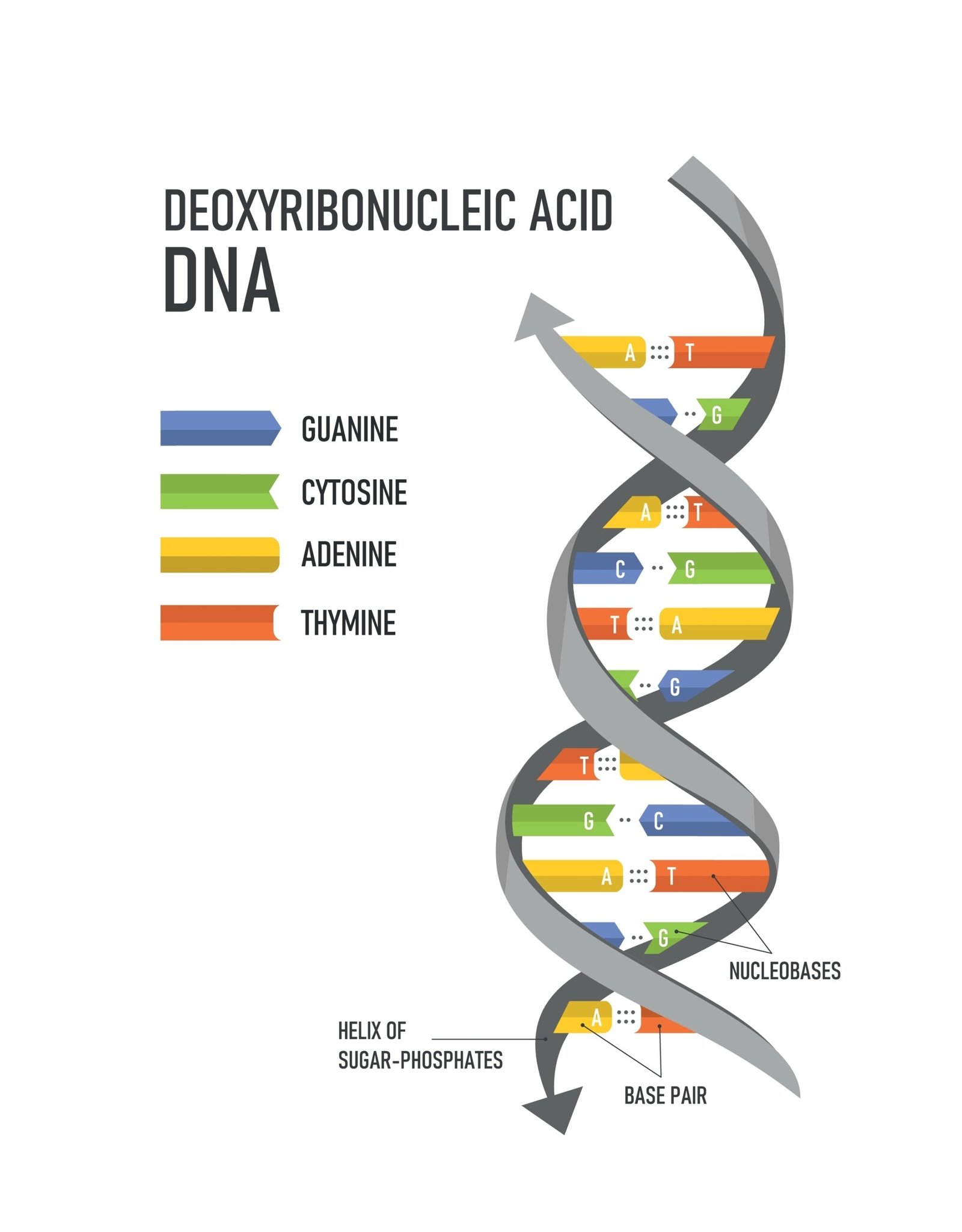
Proposed by Watson and Crick (1953).
🌀 Double helix model: two strands antiparallel, complementary base pairing (A=T, G≡C).
🔗 Chargaff’s Rule: Purine = Pyrimidine.
🔄 Hydrogen bonds: A–T (2 bonds), G–C (3 bonds).
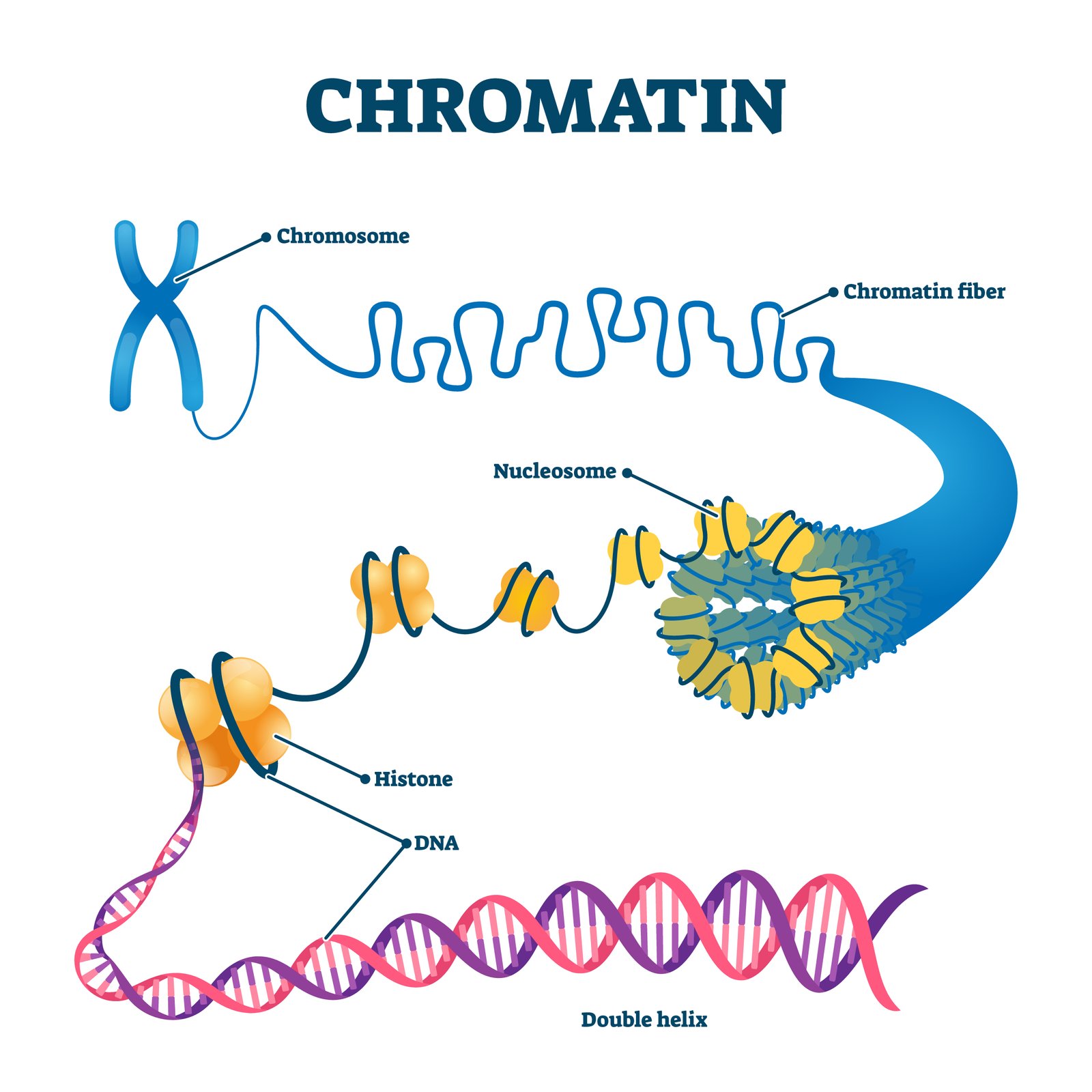
📊 Packaging: In prokaryotes → nucleoid; in eukaryotes → chromatin (DNA + histones → nucleosome “beads on a string”).
🔄 DNA Replication
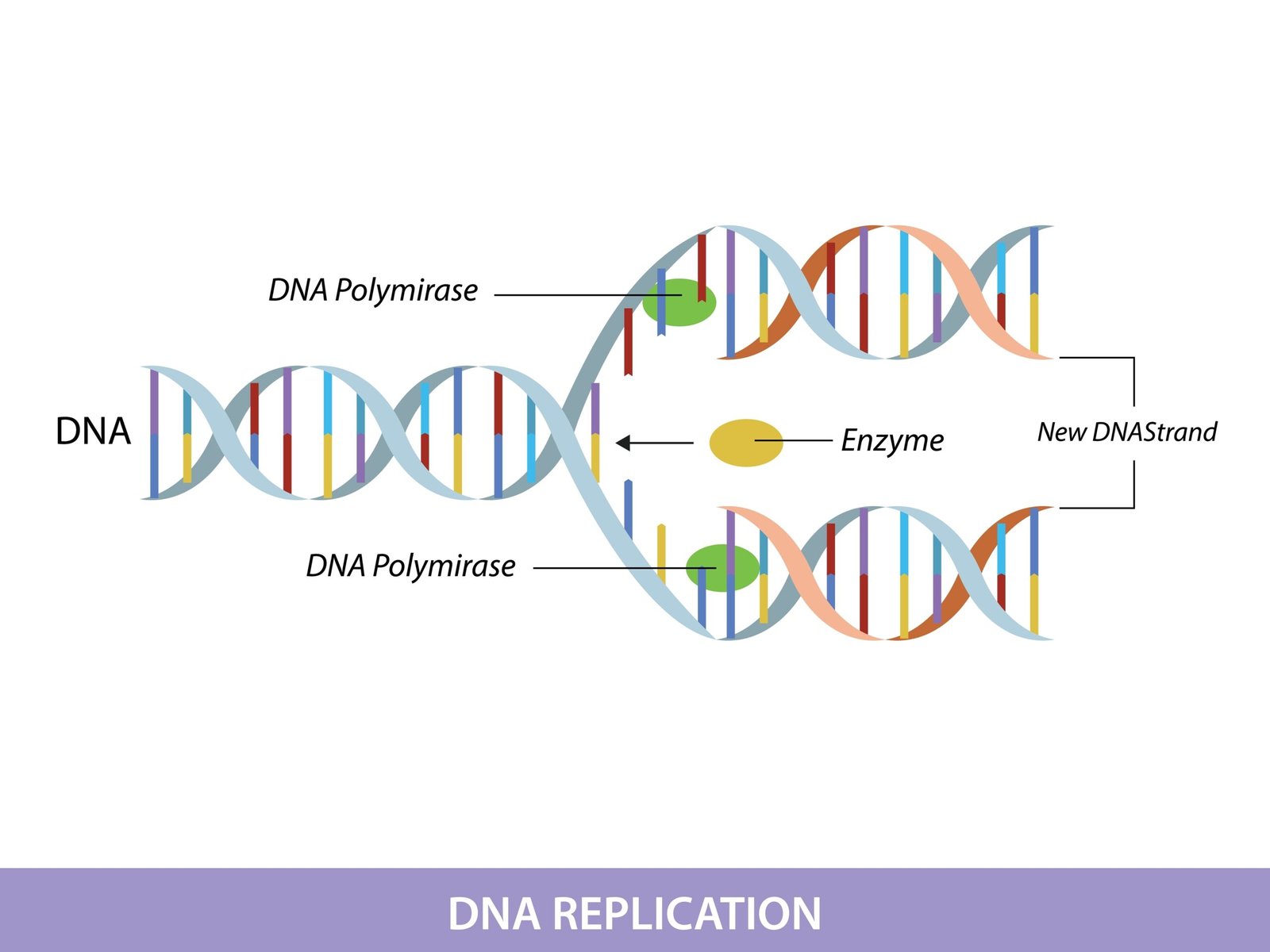
Semi-conservative (Meselson & Stahl experiment, 1958).
Steps:
Unwinding by helicase.
Primase adds RNA primer.
DNA polymerase III elongates new strand (5′→3′).
Leading strand continuous; lagging strand discontinuous → Okazaki fragments.
DNA ligase seals nicks.
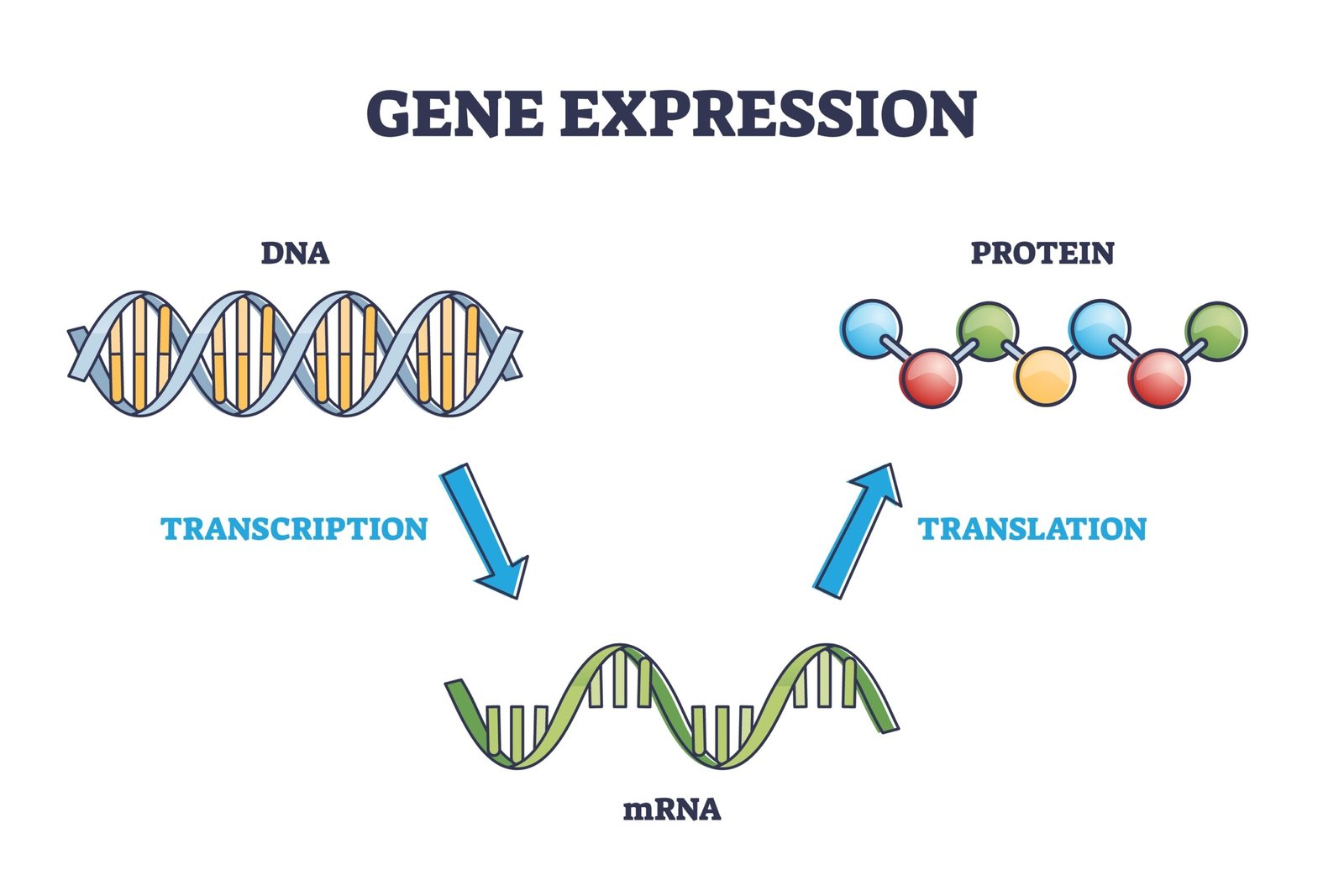
✏️ Transcription (DNA → RNA)

In prokaryotes: Entire process in cytoplasm; single RNA polymerase.
In eukaryotes: Occurs in nucleus; RNA polymerase I (rRNA), II (mRNA), III (tRNA).
Process: Initiation → Elongation → Termination.
Post-transcriptional modifications (eukaryotes):
5′ capping.
3′ poly-A tail.
Splicing (removal of introns, joining of exons).
🧪 Genetic Code
Triplet codons; 64 codons total.
Features: Universal, degenerate, non-overlapping.
AUG = Start codon (methionine).
UAA, UAG, UGA = Stop codons.
🔁 Translation (Protein Synthesis)
Ribosomes = site of translation.
Process:
Initiation → mRNA binds ribosome, tRNA brings amino acid.
Elongation → peptide bond formation.
Termination → stop codon, release factor.
🧩 Regulation of Gene Expression
Prokaryotes: Lac Operon model (Jacob & Monod).
Inducible system: presence of lactose inactivates repressor, transcription occurs.
Eukaryotes: Multiple levels (transcriptional, post-transcriptional, translational, post-translational).
🌍 Human Genome Project (HGP)
Started 1990, completed 2003.
Aim: Sequence entire human genome.
Findings: 3 billion base pairs, ~20,000–25,000 genes.
Applications: Gene therapy, DNA fingerprinting, medical diagnostics.
🧬 DNA Fingerprinting
Developed by Alec Jeffreys (1985).
Uses VNTRs (Variable Number Tandem Repeats).
Applications: Forensics, paternity testing, genetic diversity studies.
📝 Summary (~300 words)
The molecular basis of inheritance explores DNA as the genetic material. Experiments by Griffith, Avery, and Hershey–Chase established DNA’s role. The Watson–Crick double helix explains structure with base-pair complementarity. DNA replication is semi-conservative, confirmed by Meselson–Stahl. DNA replication requires helicase, primase, DNA polymerase, and ligase.
Transcription produces RNA: in prokaryotes via a single polymerase, in eukaryotes via RNA polymerase I, II, III. Eukaryotic transcripts undergo capping, tailing, and splicing. The genetic code is triplet, universal, degenerate, with AUG as start codon and UAA, UAG, UGA as stops. Translation occurs in ribosomes through initiation, elongation, and termination.
Gene expression is regulated by operons in prokaryotes, notably the lac operon. In eukaryotes, regulation is complex at multiple levels. The Human Genome Project sequenced all human genes, revolutionising medicine and diagnostics. DNA fingerprinting, based on VNTRs, has applications in forensics and identity testing.
🎯 Quick Recap
🟢 DNA → Genetic material, double helix.
🔵 Replication → Semi-conservative.
🟠 Transcription → DNA → RNA (with modifications in eukaryotes).
🔴 Translation → Protein synthesis on ribosomes.
🟡 Genetic code → Triplet, universal, degenerate.
🟣 Regulation → Lac operon, eukaryotic control.
🟤 HGP → Sequencing all human genes.
⚫ DNA fingerprinting → Forensics & identity.
————————————————————————————————————————————————————————————————————————————
QUESTIONS FROM TEXTBOOK
🔹 Q1. Group the following as nitrogenous bases and nucleosides: Adenine, Cytidine, Thymine, Guanosine, Uracil and Cytosine.
✔️ Answer:
Nitrogenous bases: Adenine, Thymine, Uracil, Cytosine.
Nucleosides (base + sugar): Cytidine, Guanosine.
🔹 Q2. If a double stranded DNA has 20% cytosine, calculate the percent of adenine in the DNA.
✔️ Answer (step by step):
In DNA, %C = %G.
If cytosine = 20%, guanine = 20%.
So, C + G = 40%.
Remaining = 60% → A + T.
Since A = T, %A = 30%.
🔹 Q3. If the sequence of one strand of DNA is written as follows:
5′-ATGCATGCATGCATGCATGC-3′
Write down the sequence of complementary strand in 5′→3′ direction.
✔️ Answer:
Complementary strand = 5′-GCATGCATGCATGCATGCAT-3′
🔹 Q4. If the sequence of the coding strand in a transcription unit is written as follows:
5′-ATGCATGCATGCATGCATGC-3′
Write down the sequence of mRNA.
✔️ Answer:
Coding strand = mRNA (except T replaced by U).
mRNA = 5′-AUGCAUGCAUGCAUGCAUGC-3′
🔹 Q5. Which property of DNA double helix led Watson and Crick to hypothesise semi-conservative mode of DNA replication? Explain.
✔️ Answer:
The complementary base pairing in double helix (A pairs with T, G pairs with C) suggested that each strand can act as a template for the formation of a new strand.
This led to the semi-conservative model where one parental strand is conserved and one new strand is synthesized.
🔹 Q6. Depending upon the chemical nature of the template (DNA or RNA) and the nature of nucleic acids synthesised from it (DNA or RNA), list the types of nucleic acid polymerases.
✔️ Answer:
DNA-dependent DNA polymerase → DNA synthesis from DNA template.
DNA-dependent RNA polymerase → RNA synthesis from DNA template.
RNA-dependent DNA polymerase (reverse transcriptase) → DNA synthesis from RNA template.
RNA-dependent RNA polymerase → RNA synthesis from RNA template.
🔹 Q7. How did Hershey and Chase differentiate between DNA and protein in their experiment while proving that DNA is the genetic material?
✔️ Answer:
They labeled DNA with ³²P (phosphorus isotope) and protein coat with ³⁵S (sulphur isotope) in bacteriophage.
After infection of bacteria, ³²P was found inside cells while ³⁵S remained outside.
This proved that DNA entered the bacteria and controlled heredity, not protein.
🔹 Q8. Differentiate between the following:
(a) Repetitive DNA and Satellite DNA
✔️ Answer:
Repetitive DNA: Occurs many times, dispersed throughout genome, no specific function.
Satellite DNA: Highly repetitive, present in tandem at specific regions (centromeres, telomeres), used in DNA fingerprinting.
(b) mRNA and tRNA
✔️ Answer:
mRNA: Carries genetic code from DNA to ribosome; template for protein synthesis.
tRNA: Brings amino acids to ribosome; has anticodon complementary to mRNA codon.
(c) Template strand and Coding strand
✔️ Answer:
Template strand: Used by RNA polymerase for transcription; complementary to RNA.
Coding strand: Same sequence as RNA (except T replaced by U); not transcribed.
🔹 Q9. List two essential roles of ribosome during translation.
✔️ Answer:
Provides site for mRNA and tRNA binding.
Catalyses peptide bond formation between amino acids.
🔹 Q10. In the medium where E. coli was growing, lactose was added, which induced the lac operon. Then, why does lac operon shut down some time after addition of lactose in the medium?
✔️ Answer:
Lactose acts as inducer by inactivating repressor → operon switches on.
After lactose is consumed, inducer concentration decreases → repressor binds again → transcription stops.
🔹 Q11. Explain (in one or two lines) the function of the following:
(a) Promoter
✔️ Provides binding site for RNA polymerase, initiates transcription.
(b) tRNA
✔️ Transfers specific amino acids to ribosome during translation, decodes mRNA codons.
(c) Exons
✔️ Coding sequences of eukaryotic genes that remain in mature mRNA after splicing.
🔹 Q12. Why is the Human Genome Project called a mega project?
✔️ Answer:
Enormous scale: sequencing 3 × 10^9 base pairs.
Required high technology, large funds, collaboration of many scientists globally.
🔹 Q13. What is DNA fingerprinting? Mention its application.
✔️ Answer:
DNA fingerprinting: Technique to identify individuals based on unique DNA patterns (VNTRs).
Applications: Forensic analysis, paternity disputes, population genetics, crime detection.
🔹 Q14. Briefly describe the following:
(a) Transcription
✔️ Process of synthesizing RNA from DNA template using RNA polymerase.
(b) Polymorphism
✔️ Genetic variation at a locus within a population, basis of evolution and DNA fingerprinting.
(c) Translation
✔️ Process of protein synthesis from mRNA using ribosomes, tRNA, and amino acids.
(d) Bioinformatics
✔️ Application of computational tools to manage and analyze biological data, especially genome sequences.
————————————————————————————————————————————————————————————————————————————
OTHER IMPORTANT QUESTIONS FOR EXAMS
(CBSE MODEL QUESTION PAPER)
ESPECIALLY MADE FROM THIS CHAPTER ONLY
🔹 Q1. (MCQ) 🦠 Hershey–Chase used labeled phages to prove DNA is genetic material. The correct isotope pair was:
🔵 (A) ¹⁵N and ¹⁴N
🟢 (B) ³²P and ³⁵S
🟠 (C) ³H and ¹⁴C
🔴 (D) ⁶⁰Co and ³²S
✔️ Answer: (B) ³²P and ³⁵S
🔹 Q2. (MCQ) 🧬 The feature degeneracy of the genetic code means:
🔵 (A) One codon codes for multiple amino acids
🟢 (B) Multiple codons can code for one amino acid
🟠 (C) Codons overlap during reading
🔴 (D) Codons have punctuation marks
✔️ Answer: (B) Multiple codons can code for one amino acid
🔹 Q3. (MCQ) 🧪 The enzyme that seals nicks between Okazaki fragments is:
🔵 (A) DNA polymerase I
🟢 (B) DNA ligase
🟠 (C) RNA primase
🔴 (D) Topoisomerase
✔️ Answer: (B) DNA ligase
🔹 Q4. (MCQ) 🧫 In eukaryotes, RNA polymerase II synthesizes:
🔵 (A) rRNA
🟢 (B) mRNA
🟠 (C) tRNA
🔴 (D) 5S rRNA only
✔️ Answer: (B) mRNA
🔹 Q5. (MCQ) 🌱 The repeating structural unit of chromatin is:
🔵 (A) Solenoid
🟢 (B) Nucleosome
🟠 (C) Looped domain
🔴 (D) Chromatid
✔️ Answer: (B) Nucleosome
🔹 Q6. (MCQ) 🧬 The coding strand of a gene has the same sequence as:
🔵 (A) tRNA (with T instead of U)
🟢 (B) mRNA (with T replaced by U)
🟠 (C) Template strand
🔴 (D) rRNA (with T replaced by U)
✔️ Answer: (B) mRNA (with T replaced by U)
🔹 Q7. (MCQ) 🧪 The start codon in most organisms is:
🔵 (A) UAA
🟢 (B) AUG
🟠 (C) UAG
🔴 (D) UGA
✔️ Answer: (B) AUG
🔹 Q8. (MCQ) 🦋 Lac operon is inducible because the inducer is:
🔵 (A) Lactose/allolactose that inactivates the repressor
🟢 (B) Glucose that activates CAP
🟠 (C) Tryptophan that activates repressor
🔴 (D) IPTG that blocks promoter
✔️ Answer: (A) Lactose/allolactose that inactivates the repressor
🧠 Assertion–Reason format (for Q9–Q10):
1 = A true, R true, R explains A; 2 = A true, R true, R not explanation; 3 = A true, R false; 4 = A false, R true.
🔹 Q9. (A–R MCQ) 🧬 Assertion (A): In eukaryotes, primary transcript (hnRNA) undergoes capping, tailing, and splicing.
Reason (R): Eukaryotic genes are often interrupted by introns that must be removed.
🔵 (1)
🟢 (2)
🟠 (3)
🔴 (4)
✔️ Answer: (1)
🔹 Q10. (A–R MCQ) 🧫 Assertion (A): DNA replication proceeds 5’→3′ on both new strands.
Reason (R): DNA polymerase can extend only from a free 3′-OH end.
🔵 (1)
🟢 (2)
🟠 (3)
🔴 (4)
✔️ Answer: (1)
🟡 Very Short Answer (Q11–Q18, 1 mark each)
🔹 Q11. 🧪 Define Okazaki fragments.
✔️ Answer: Short DNA segments synthesized discontinuously on the lagging strand during replication.
🔹 Q12. 🧬 State Chargaff’s rule.
✔️ Answer: In dsDNA, [A] = [T] and [G] = [C]; purines equal pyrimidines.
🔹 Q13. 🧫 Name the inducer of lac operon in vivo.
✔️ Answer: Allolactose (isomer of lactose).
🔹 Q14. 🦠 Full form of VNTR used in DNA fingerprinting.
✔️ Answer: Variable Number Tandem Repeats.
🔹 Q15. 🌱 Two stop codons.
✔️ Answer: UAA, UAG (also UGA).
🔹 Q16. 🧬 Name the experiment that proved semi-conservative replication.
✔️ Answer: Meselson–Stahl experiment.
🔹 Q17. 🧪 Enzyme that removes RNA primers in prokaryotes.
✔️ Answer: DNA polymerase I (5’→3′ exonuclease).
🔹 Q18. 🧫 Type of chromatin that is transcriptionally active.
✔️ Answer: Euchromatin.
🟠 Short Answer (Q19–Q27 carry 2–3 marks each) — Part 1 includes Q19–Q20
🔹 Q19. 🧬 Calculate base composition: In a dsDNA, adenine = 36%. Find % of T, G, C.
✔️ Answer (step-by-step):
A = 36% → T = 36%.
A + T = 36 + 36 = 72%.
Therefore G + C = 100 − 72 = 28%.
Hence G = 14% and C = 14%.
🔹 Q20. 🧪 The coding strand of a gene is 5′-ATG GAA TGC TTT GGA-3′. Write the template strand and mRNA sequence.
✔️ Answer (step-by-step):
Coding strand (5’→3′) = 5′-ATG GAA TGC TTT GGA-3′.
Template strand is complementary and antiparallel → 3′-TAC CTT ACG AAA CCT-5′ (or written 5′-TTC CAA A GC AAG GTA? No—keep 3’→5′ form).
mRNA has same sequence as coding strand with U for T → 5′-AUG GAA UGC UUU GGA-3′.
🔹 Q21. 🦠 Write any three features of the genetic code.
✔️ Answer:
Triplet: Three bases code for one amino acid.
Universal: Same in almost all organisms.
Degenerate: More than one codon can code for a single amino acid.
🔹 Q22. 🧬 Differentiate between replication in prokaryotes and eukaryotes (two points).
✔️ Answer:
Prokaryotes: Single origin of replication, faster.
Eukaryotes: Multiple origins of replication, slower.
🔹 Q23. 🧪 Explain splicing, capping, and tailing in eukaryotic mRNA.
✔️ Answer:
Splicing: Removal of introns, joining of exons.
Capping: Addition of methyl guanosine triphosphate at 5′ end.
Tailing: Addition of poly-A tail at 3′ end.
🔹 Q24. 🌱 What is DNA fingerprinting? Mention two applications.
✔️ Answer:
Technique to identify individuals based on unique DNA patterns (VNTRs).
Applications: Forensic identification, paternity disputes.
🔹 Q25. 🧫 What is the role of the repressor protein in lac operon?
✔️ Answer:
Repressor binds to operator region and prevents transcription in absence of lactose.
In presence of lactose, allolactose binds repressor → repressor inactivated → transcription occurs.
🔹 Q26. 🧬 Define polymorphism. How is it important in DNA fingerprinting?
✔️ Answer:
Polymorphism: Genetic variation at a locus present in >1% population.
Provides unique DNA profiles used in DNA fingerprinting.
🔹 Q27. 🧪 What is the role of tRNA during translation?
✔️ Answer:
Carries specific amino acids to ribosome.
Anticodon of tRNA pairs with codon of mRNA to ensure correct amino acid placement.
🟣 Long Answer Questions (5 marks each)
🔹 Q28. 🧬 Explain the Meselson–Stahl experiment proving semi-conservative replication.
✔️ Answer:
E. coli grown in ¹⁵N medium → DNA heavy.
Transferred to ¹⁴N medium → after 1 generation, hybrid DNA.
After 2 generations, 50% hybrid, 50% light DNA.
Proved that each DNA molecule has one parental and one new strand.
🔹 Q29. 🌿 Describe the structure of DNA as proposed by Watson and Crick.
✔️ Answer:
Double helix model.
Two antiparallel strands (5′→3′ and 3′→5′).
Bases paired: A–T (2 H-bonds), G–C (3 H-bonds).
Sugar–phosphate backbone outside, bases inside.
Uniform diameter ~2 nm, 10 base pairs per turn.
🔹 Q30. 🧪 Explain DNA replication in detail with enzymes.
✔️ Answer:
Initiation: Helicase unwinds DNA; primase adds RNA primer.
Elongation: DNA polymerase synthesizes new strands.
Leading strand: continuous.
Lagging strand: discontinuous (Okazaki fragments).
Termination: RNA primers replaced; DNA ligase joins fragments.
🔹 Q31. 🦠 Explain transcription in prokaryotes and eukaryotes.
✔️ Answer:
Prokaryotes: Single RNA polymerase binds promoter → elongation → termination by hairpin loop or rho factor.
Eukaryotes: RNA polymerase I (rRNA), II (mRNA), III (tRNA). Post-transcriptional modifications (splicing, capping, tailing).
🔹 Q32. 🧩 Explain the lac operon model of gene regulation.
✔️ Answer:
Operon consists of promoter, operator, regulator, structural genes (Z, Y, A).
Without lactose: Repressor binds operator → no transcription.
With lactose: Allolactose inactivates repressor → transcription proceeds.
Inducible system (genes expressed only when substrate is present).
🔹 Q33. 🧬 Explain the Human Genome Project (HGP) and its applications.
✔️ Answer:
Mega project (1990–2003), sequenced ~3 billion base pairs.
Found ~20,000–25,000 genes, less than expected.
Applications: Gene therapy, medical diagnostics, forensics, evolution studies.
————————————————————————————————————————————————————————————————————————————
